APGI BioResource: Introduction & Availability
APGI BioResource
From 2016 the APGI BioResource is proudly supported by a grant from PanKind, Pancreatic Cancer Australia (previously The Avner Pancreatic Cancer Foundation)
Our tissue and data resource contains fresh frozen tissue, formalin fixed paraffin embedded (FFPE) tissue, blood, blood products, tissue microarrays (TMAs), patient-derived xenografts (PDX) and cell lines (PDCL) across a spectrum of pancreatic diseases. This resource has proven to be of consistent high-quality, and has been the foundation of several landmark publications in Pancreatic Cancer (Nature 2012, Nature 2013, Nature 2016).
The APGI BioResource of biospecimens and clincial and genomic data is one of the largest and most comprehensive in the world. The APGI houses a dedicated team to coordinate sample collection, management, processing and distribution based at the Garvan Institute of Medical Research in Sydney.
We currently have over 4,000 pancreatic cancer patients in our database, with a range of prospectively collected and archived biospecimens. Every biospecimen is coupled with detailed clinico-pathological data, including past medical history, treatment data and detailed disease outcomes. Our resource is updated regularly and maintained by a dedicated team. We have personal connection to most of our clincal teams and follow our patients closely through their disease journey and beyond. This makes us one of the largest resources of pancreatic biological material in the world, and our samples have been used to support over 200 research studies globally.
Please read the APGI BioSpecimen and Data Access Policy, which outlines requirements for distribution and use of our resource. For further information about how you can use the BioResource for your research, please contact us at research@pancreaticcancer.net.au.

BioSpecimen Summary and Availability
Quality Standards
We have been collecting, processing, analysing and storing biospecimens since 2006. Through this process we have developed state-of-the-science biospecimen resource practices, based on the guidelines from international bodies such as the NCI Best Practices for Biospecimen Resources. These practices and standard operating procedures outline the operational, technical, ethical, legal and policy best practices for the APGI BioResource, and are specific to the mission and scientific needs of the pancreatic cancer translational research community. Our best practices and detailed collection, processing and distribution protocols promote biospecimen and data quality and support adherence to ethical and legal requirements.
Tissue MicroArray (TMA)
The Avner APGI BioResource includes one of the world’s largest and most comprehensively annotated FFPE pancreatic tumour tissue cohort in micro-array format. This retrospectively acquired Australian surgical cohort spans over a decade, and is especially designed for rapid biomarker discovery studies. Prospectively collected FFPE tissue is currently being used to construct further TMAs. Pancreatic tumour types include pancreatic ductal adenocarcinomas, ampullary carcinomas, intraductal papillary mucinous neoplasms, pancreatic endocrine tumours and solid pseudo papillary tumours.
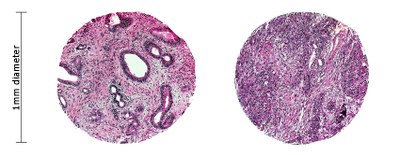
← This diagram shows two representative tissue micro-array cores viewed under light microscopy at medium to high magnification. Our arrays typically contain up to 96 cores per slide. On the left is an example of a moderately differentiated pancreatic ductal adenocarcinoma, while on the right is a poorly differentiated example.
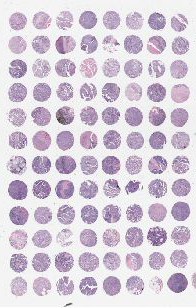
Each TMA* contains triplicates of each core as well as controls. TMA maps* displaying the location and minimal relevant information of each core will be provided once samples are distributed.
*Excludes Test Arrays
| BioSpecimen Type | Patient count | TMA count | Restricted | Cohort Description |
|---|---|---|---|---|
| Pancreatic Ductal Adenocarcinoma (PDAC) Test Array | 20 cases | 1 TMA | No | For optimizing purposes |
| Non-neoplastic Pancreas Test Array | 20 cases | 1 TMA | No | For optimizing purposes |
| Pancreatic Ductal Adenocarcinoma (PDAC) Training Set Arrays | 96 cases | 3 TMAs | No | See Pinho et al 2016 |
| Pancreatic Ductal Adenocarcinoma (PDAC) Validation Set Arrays | 352 cases | 11 TMAs | No | See Biankin et al 2009 |
| Ampullary Carcinoma Set Arrays | 96 cases | 3 TMAs | No | See Chang et al 2013 |
| Intraductal Papillary Mucinous Neoplasm (IPMN) Set Arrays | 56 cases | 2 TMAs | No | See Biankin et al 2002 |
| Solid Pseudopapillary Tumour (SPT) Set Array | 32 cases | 1 TMA | No | |
| Pancreatic Intraductal Neoplasm (PanIN) Array | 32 cases | 1 TMA | No | |
| Chronic Pancreatitis Set Arrays | 64 cases | 2 TMAs | No | |
| Pancreatic Neuroendocrine (PNET) Set Arrays | 64 cases | 2 TMAs | No | |
| PDAC International Cancer Genome Consortium (ICGC) Set Arrays | 232 cases | 8 TMAs | Yes | See Biankin et al 2012 |
| PDAC International Cancer Genome Consortium (ICGC) Refined Set Arrays | 39 cases | 2 TMAs | No | See Waddell et al 2015 |
| PNET International Cancer Genome Consortium (ICGC) Arrays | 46 Cases | 2 TMAs | No | See Scarpa et al 2017 |
Formalin Fixed Paraffin Embedded (FFPE) Tissue
A wide range of FFPE tissue is available from our BioResource. We do not provide whole FFPE tissue blocks but can arrange whole sections (either on slides or in cigars) or cores for downstream processes such as nucleic acid extraction or staining.
Many of our prospectively collected FFPE tissue have matched snap frozen and blood component material, accompanied by a rich clinicopathological dataset.
Fresh Tissue
Surgically resected tissue of various histological types, including limited metastases, have been collected and snap frozen in liquid nitrogen. In some cases, tissue in DMSO may be available as well as patient-derived xenograft (PDX) tissue.
Cell Lines
Patient-derived cell lines (PDCL’s) have been created on 18 patients. Some patients have multiple cell lines grown under different environments. If you are requesting PDCL’s only, without any clinical information, the approval process may be expedited. Read more about accessing PDCL’s here.
Blood
Received whole blood is centrifuged and divided into resulting blood components – plasma, buffy coat, red blood cells, and serum (where applicable) – and snap frozen in liquid nitrogen.
Whole blood may also be available, sometimes in the form of PAXgene DNA Blood Tubes.
Nucleic Acid Analytes
DNA and RNA analytes extracted from various BioSpecimen types are stored in buffer at -80°C, with accompanying BioSpecimen data (concentration, volume, A260/280, etc) recorded for later aliquot creation.
High Risk Cohorts
As part of the Australian familial Pancreatic Cancer Cohort (AFPaCC) 2011-2016 we have accrued a cohort of over 200 families with a history of pancreatic cancer through a dedicated registry. The cohort includes blood, tissue samples (mostly FFPE) and detailed clinical, pathological and family history information on families deemed High risk. This is cohort of patients and samples are available for the research community to access and would be a valuable adjunct to genetic or clinical studies on familial cancer. Further details are available on our Familial pages.
Images
Scanned H&E images are available for the ICGC sequenced fresh tissue BioSpecimens as TIFF (thumbnail) files with a dimension of about 1024×768 pixels.
Full resolution SVS files may be available upon request however sufficient time and storage (approximately 3 GB per image) needs to be made available.
Clinicopathological Data
The APGI collects and stores data on over 500 datapoints for each consented participant. This data varies across multiple sections and is one of the most complete in the world on pancreatic cancer patients:
- Demographics
- Clinical Presentation
- Risk Factors
- Comorbidities
- Family History
- BioAssay
- Surgery
- Medications
- Pathology
- Chemotherapy
- Radiotherapy
- Outcome
- Recurrence
- BioSpecimens
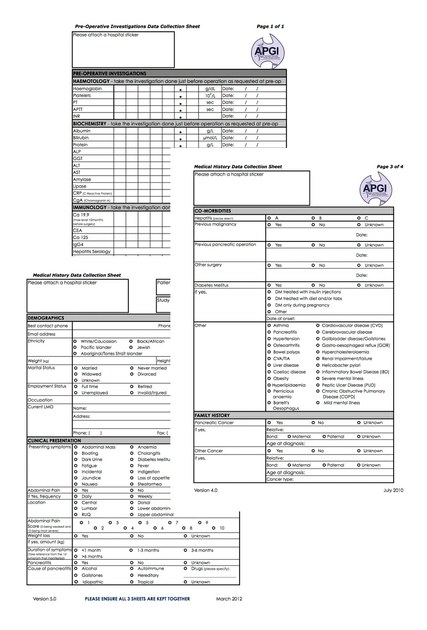
Data Collection Sheets are completed by APGI staff to gather data from multiple sites throughout Australia including GP or specialist rooms and hospital medical records. This information is then entered into CanSto.

↓ Some of the APGI’s Data Collection Sheets are displayed in the image below. These user-friendly sheets guide staff on datapoints to be collected whilst off-site for later electronic input.
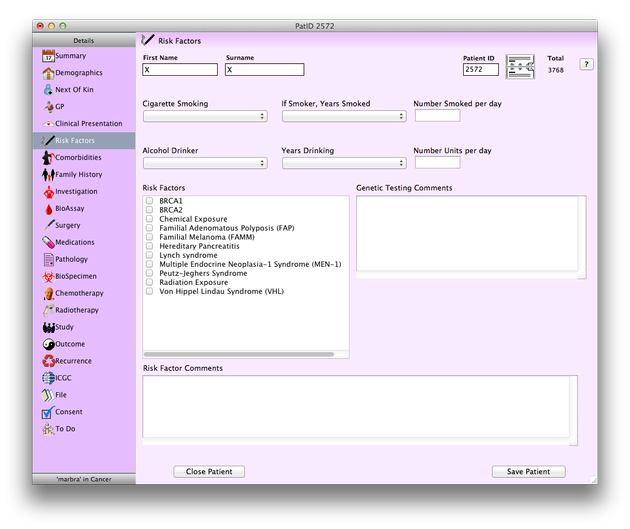
↓ This screenshot is from the APGI’s custom database CanSto Pancreas. The panel on the left displays the multiple sections within the database where all participant data is recorded. The Risk Factors section is selected in the panel here therefore on the right the Risk Factors section appears. Data can be entered via the various available options i.e. selecting from drop-down or checkbox menus or typing data directly into text fields.